Calibrate ensemble¶
You can jointly calibrate multiple models as an ensemble. In an ensemble calibration:
- Each member model is independently calibrated against the same time-series dataset representing historical observations.
- The models are recalibrated as a single ensemble, with loss or prediction error calculated as a linearly weighted sum of the different model outputs and dataset features.
The result is a joint set of calibrated model configurations that may generate predictions with lower error than than individual models or their independently calibrated configurations.
Ensemble calibration essentially allows a search for configuration solutions where the member models can specialize to different features or time periods of the given dataset. This is analogous to a high-performing random forest regressor which can be constructed from multiple weak-learning decision trees.
Calibrate ensemble is powered by the ensemble_calibrate function of the PyCIEMSS package. An example of how it can be used programmatically can be found in this Jupyter notebook.
Tip
You can quickly create an ensemble calibration using the Calibrate an ensemble model workflow template.
Calibrate ensemble operator¶
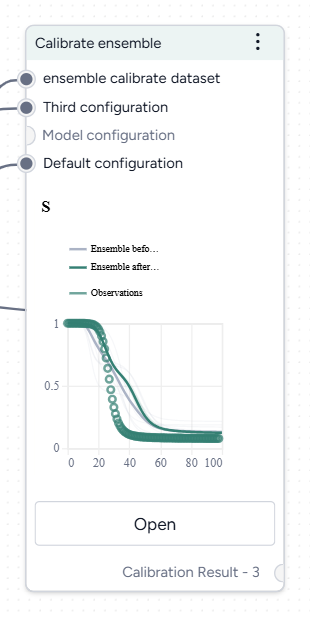
In a workflow, the Calibrate ensemble operator takes two or more model configurations and a dataset as inputs. It outputs a calibrated dataset.
Once you've completed the calibration, the thumbnail preview shows the calibrated ensemble variables over time.
-
Inputs
- Two or more model configurations
- Dataset (with timepoints)
-
Outputs
Calibrated dataset
Add a Calibrate ensemble operator to a workflow
-
Do one of the following actions:
- On an operator that outputs a model configuration, click Link > Calibrate ensemble.
- Right-click anywhere on the workflow graph, select Simulation > Calibrate ensemble, and then connect two or more model configurations and a dataset to the Calibrate ensemble input.
Calibrate ensemble¶
The Calibrate ensemble operator allows you to define how to:
- Map your model configurations and dataset.
- Assign your level of confidence in each model.
- Choose how to run the calibration.
Open a Calibrate ensemble operator
- Make sure you've connected two or more model configurations and a dataset to the Calibrate ensemble operator.
- Click Open.
Map dataset columns and model variables¶
To begin, create a mapping between the calibration data and the model configurations by selecting features and corresponding outcomes that are represented by each model.
Example
Map dataset columns current number of hospitalized cases and cumulative number of deaths to threatened (T) and extinct (E) populations in a SIDARTHE model or hospitalizations (H) and deaths (D) in a SEIQHRD-type model
It's not necessary to map between every dataset feature to a set of model outcomes. For example, susceptible or exposed populations in SEIR-type models do not correspond to readily observed or reported values.
Tip
High-quality case, death, and hospitalization data for COVID-19 is available on the COVID-19 Forecast Hub GitHub repository. If a model selected for ensemble calibration does not have a state variable that corresponds well to these observations, add an observable using Edit model notebook.
Tip
State variables in compartmental models usually represent prevalence of disease conditions. Literature and data repositories often provide only daily incident or cumulative estimates. Use the Transform dataset operator to convert from incidence to prevalence values.
You can also use the Edit model operator to add new "controlled production" transitions and create new state variables that are cumulative equivalents.
For example, define a cumulative equivalent (Icum) of the number of infected individuals (I) for a SIR-type model by adding a controlled production where the:
- Outcome is
Icum. - Controller is
I. - Rate law is
I.
The resulting equation in this case should be d Icum(t) / dt = I(t) or Icum(t) = int I(t) dt.
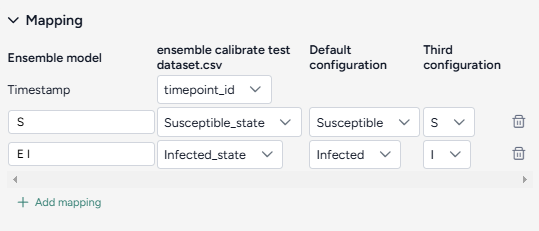
Create a mapping between the calibration data and the model configurations
- Select the Timestamp column from the dataset.
-
For each variable of interest:
- Click Add mapping.
- Enter a unique name for the variable in the ensemble model.
- Select the corresponding state from each of the model configurations.
Assign model weights¶
Model weights are the parameters used to linearly sum the model outcomes in the ensemble. They represent how much each model contributes to the ensemble.
Because Calibrate ensemble is Bayesian inference process, the model weights are not single values but are represented by a Dirichlet distribution of order K, where K is the number of models in the ensemble. This distribution has parameters a_1, a_2, ..., a_K that control where and how much mass is concentrated at different possible combinations of normalized model weights (w_1, w_2, ..., w_k).
In the interface, the Dirichlet alpha parameters a_k are exposed as Relative uncertainty. Selecting from 0 to 10 allows you to express your confidence in each of the models and find an ensemble solution that potentially produces better predictions.
Assign model weights
- For each model, select a value from
1(low) to10(high) using the Relative certainty dropdown.
- If you don't know which models are better and should have more weight, assign equal medium weights (
5). This is equivalent to initially assuming a flat Dirichlet distribution where all model weight combinations are equally probable. This is the default and recommended option. - Assign a high weight (
10) to one or more models that you believe produce the best predictions. Calibrate ensemble searches for solutions that prioritize the contribution of these models. - Assign a low weight (
1) to models that you believe are less reliable. Their contributions are reduced in the ensemble calculation.
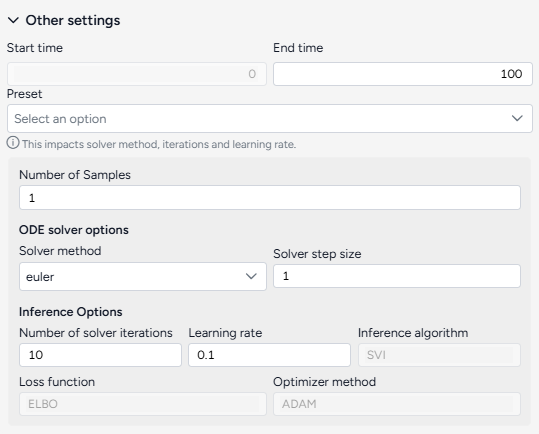
Configure the run settings¶
The Other settings control the time period of interest and behavior of the underlying ODE solver and optimizer. You can select a preset (Fast or Normal) or adjust these settings individually to balance between run time and precision.
Quickly configure the run settings
- Choose the Start time and End time to specify the timepoints to be simulated.
- Select a Preset, Fast or Normal.
Advanced settings
Using the following advanced settings to control how fast or precise the calibration should be:
- Number of samples: Number of point estimates to be made on the posterior distribution (model parameters and weights) for the pre and post-calibration simulations.
- ODE solver options determine the approach for solving the equations of the model during calibration:
- Solver method: dopri5 returns the best results while euler is faster but less accurate (more details).
- Solver step size: Size of the time interval used to integrate the solution. Larger values are faster but less accurate (only needed by the euler method).
- Inference options control the behavior of the inference algorithm that minimizes the calibration error:
- Number of solver iterations: The number of calibration steps.
- Learning rate: Step size for updating parameter values during calibration.
- Inference algorithm: Stochastic Variational Inference (SVI), which estimates the parameters probabilistically.
- Loss function: Evidence Lower Bound (ELBO), which guides parameter updates by balancing data fit and model complexity.
- Optimizer method: algorithm for updating parameter values, ADAM by default.
Tip
Consider using minimum settings - such as the end time at 3, the number of samples at 1, and the solver method at euler - to check whether the calibration can run to completion with the given mapping.
Create and save the calibrated dataset¶
Once you've configured the settings, you can run the operator to generate a new calibrated dataset. The new dataset becomes a temporary output for the Calibrate ensemble operator. You can connect it to other operators in the same workflow.
Create a new calibrated dataset
- Click Run.
Choose a different output for the Calibrate ensemble operator
- Use the Select an output dropdown.
Understand the results¶
Ensemble calibration results are presented as a series of charts that show:
Access the Output settings
Additional settings for the various chart types are available in the Output settings panel.
- Click Expand to expand the Output settings.
Save a chart for use outside Terarium
You can save Calibrate ensemble charts for use outside of Terarium. Download charts as images that you can share or include in reports, or access structured JSON that you can edit with Vega .
- Click and then choose one of the following options:
- Save as SVG
- Save as PNG
- View source (Vega-Lite JSON)
- View compiled Vega (JSON)
- Open in Vega Editor
Add annotations that call out key values and timesteps
To highlight notable findings, you can manually add annotations that label plotted values at key timesteps on loss and ensemble variable charts.
- Click anywhere on the chart to add a callout.
- To add more callouts without clearing the first one, hold down Shift and click a new area of the chart.
Note
Ensemble variable charts also support AI-generated annotations.
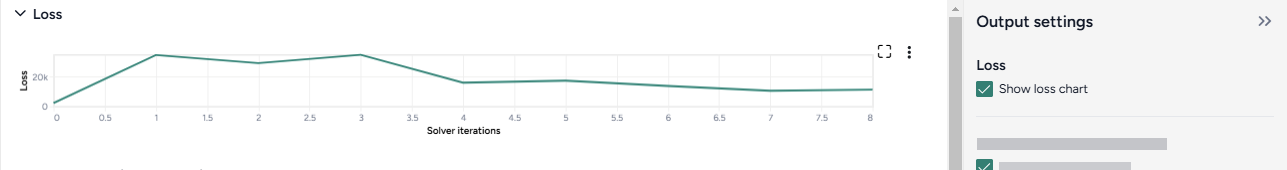
Loss¶
The loss chart shows the error between the ensemble model outputs and the calibration data. A monotonic and exponential decrease in loss values indicates convergence and a successful calibration.
Show or hide the loss chart
- Select or clear Show loss chart.
Tip
If the loss doesn't decrease to a plateau, the calibration algorithm may be struggling to converge on a solution. Failure to converge could indicate that the model outputs are not good matches to the calibration data. Consider reducing the complexity of the problem by:
- Adjusting the values of the input model configurations to approximate a solution.
- Removing the larger or least trustworthy model configurations.
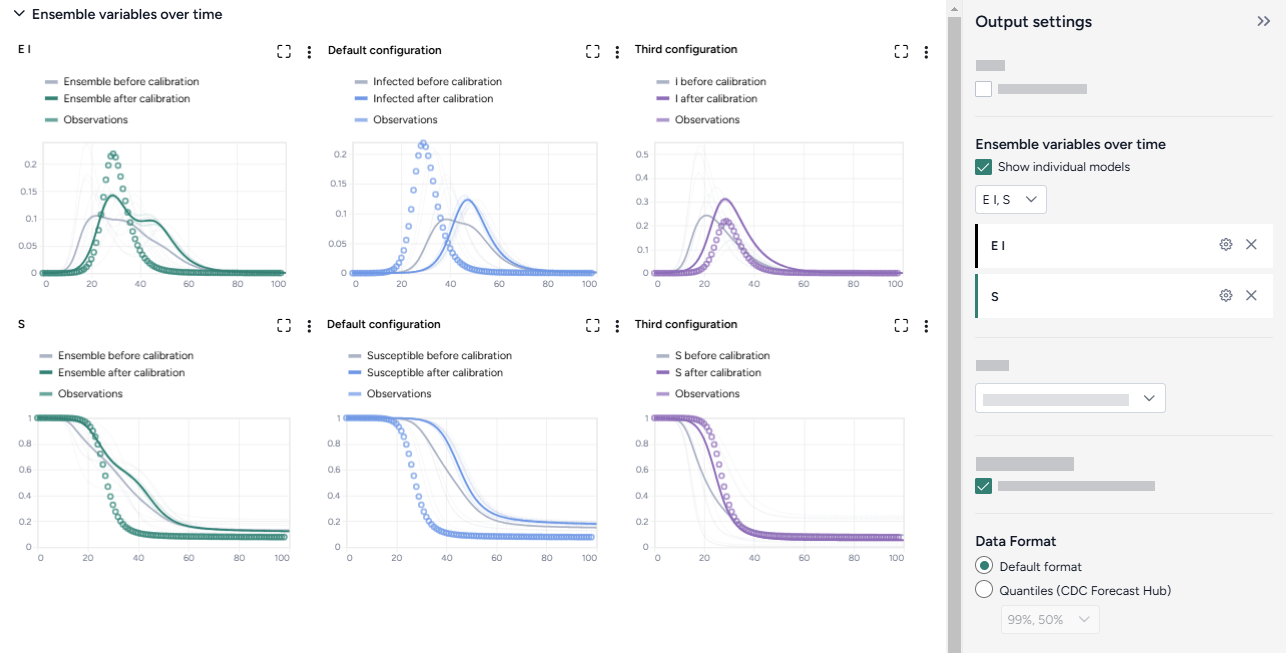
Ensemble variables over time¶
To aid visual validation, the Ensemble variables over time charts compare the effects of calibration for each state variable and the historical data.
- The grey line represents the model before calibration.
- The colored line represents the model after calibration.
- Circles represent observations from the historical data.
The Output settings panel has several options that let you customize the scale and display of the charts. You can also use an AI assistant to generate chart annotations that highlight key data points.
Choose which model states to plot
- Select the model states from the dropdown list.
Show charts for each model configuration
You can view individual charts for all the models that contribute to the ensemble model as small multiples.
- Click Show individual models.
Change the data format
The data format controls how the ensemble variable charts are drawn.
- Choose the Data format:
- Default: Include the historical observations in the plot.
- Quantiles (CDC Forecast Hub): Omit the historical observations and draw filled shapes to represent quantiles ranging from 50$ndash;90%.
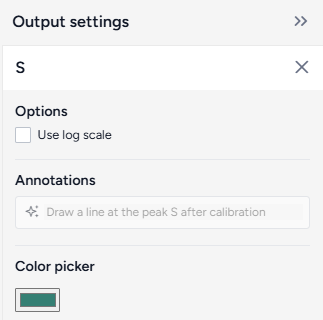
Additional chart settings are available for each of the state variables.
Access additional chart settings
- Click Options .
Change the chart scale
By default, ensemble variable charts are shown in linear scale. You can switch to log scale to view large ranges, exponential trends, and improve visibility of small variations.
- Select or clear Use log scale.
Change variable colors
You can change the color of any model state to make your charts easier to read.
- Click the color picker and choose a new color from the palette or use the eye dropper to select a color shown on your screen.
Prompt an AI assistant to add chart annotations
You can prompt an AI assistant to automatically create annotations on the ensemble variable charts. Annotations are labelled or unlabelled lines that mark specific timestamps or peak values. Examples of AI-assisted annotations are listed below.
-
Describe the annotations you want to add and press Enter.
Draw a vertical line at day 100Draw a line at the peak S after calibrationDraw a horizontal line at the peak of default configuration Susceptible after calibration. Label it as "important"Draw a vertical line at x is 10. Don't add the labelDraw a line at x = 40 only for ensemble after calibration
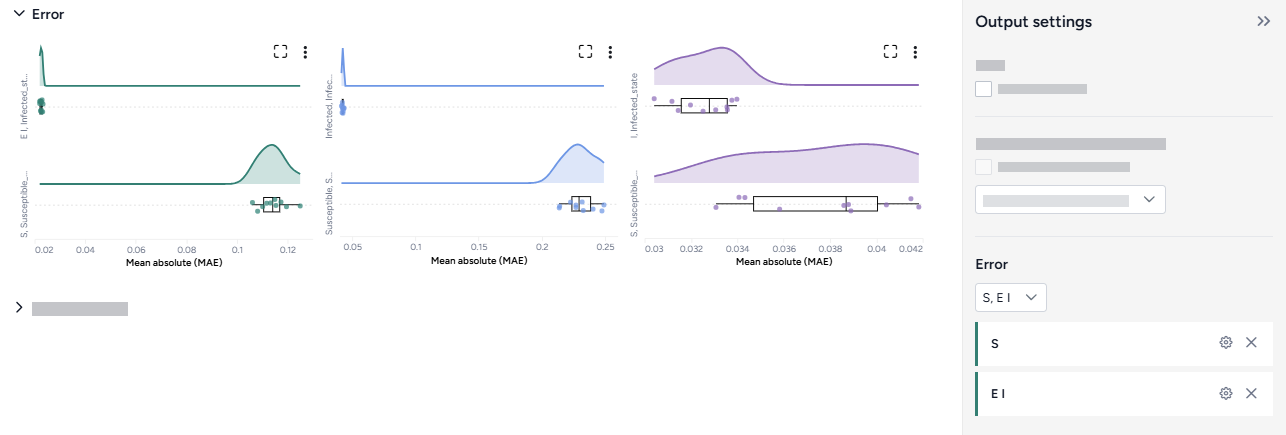
Error¶
The error plots show the mean absolute error (MAE) for each model and variable of interest.
Change the chart scale
By default, error charts are shown in linear scale. You can switch to log scale to view large ranges, exponential trends, and improve visibility of small variations.
- Click Options .
- Select or clear Use log scale.
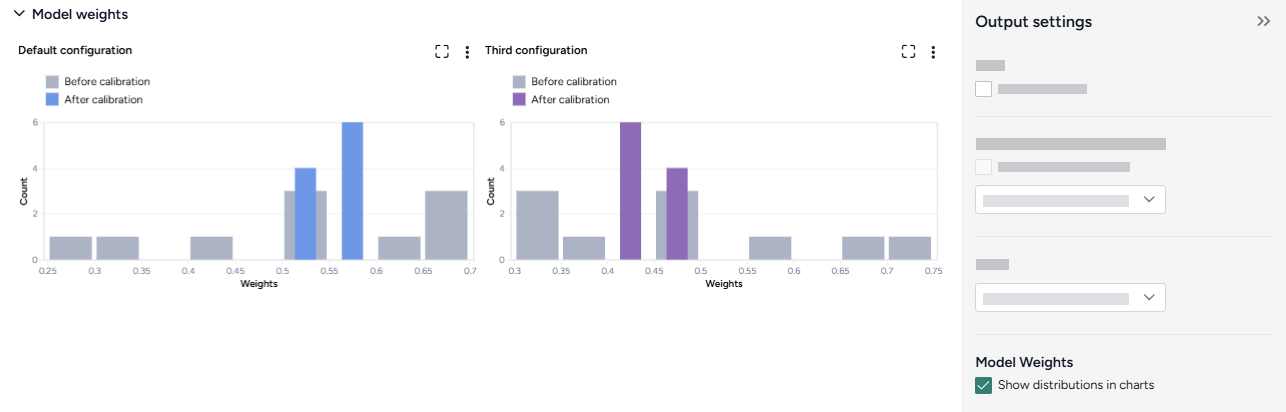
Model weights¶
The model weights charts display one-dimensional projections of the Dirichlet distribution of the weights for each model, before and after the calibration.
Generally, a good calibration takes a broad distribution (a low-certainty prior) and returns a narrow distribution (a high-certainty posterior, conditioned on the calibration data).
To show or hide the model weights charts
- Select or clear Show distributions in charts.
Troubleshooting¶
Recommended run settings¶
It's recommended you run calibrations using the dopri5 Solver method.
Calibrate each model first¶
Try calibrating each model to your dataset independently.
Relative certainty¶
Set the Relative certainty in the Model weights to 1 for each model. When using this setting, proceed slowly and cautiously. To include a preference for one model over the others, start by increasing its Relative certainty to 2, then 3, and so on.
Uncertainty and number of samples¶
If your models have no uncertainty in parameter values, only one sample is needed. Change Number of samples to 1 (the default is set to 100).
Simulation length and number of samples¶
If you plan to simulate your calibrated ensemble model for a long time or with a large number of samples (for example, End time or Number of samples > 100), set them to a lower value (10 or 20) first and run a check for errors.
Error messages¶
PyCIEMSS error messages should offer guidance on how to proceed. Error messages from Pyro or torchdiffeq may be less clear.
If you see a messages referencing Cholesky factorization (including The factorization could not be completed because the input is not positive-definite) or AssertionError with underflow in dt 0.0:
- If you successfully calibrated each model independently, check that the models and the dataset are on the same scale. Errors are likely if:
- One model assumes a population of 10 million, while another (or the dataset) has a population of 1,000.
- One model (or the dataset) is normalized to a population of one while the others are not.
- Make sure your initial conditions are similar to each other and consistent with your dataset. They don't need to be an exact match, but errors are likely if they are too far off.